- en Change Region
- Global Site
Nikon Instruments Inc. | Americas
- Home
- Products
- Confocal and Multiphoton Microscopes
- AX / AX R with NSPARC
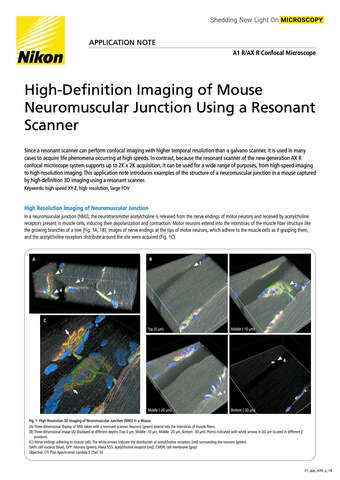
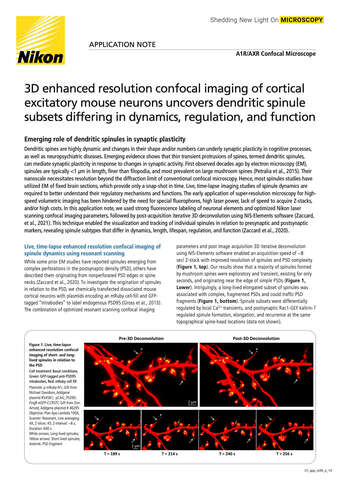
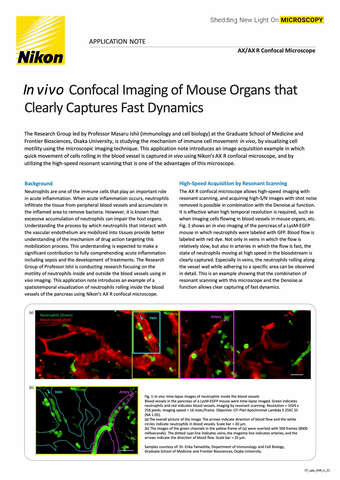
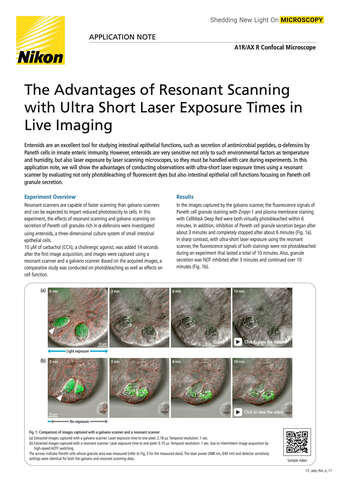
Glial cell surrounded by axons in a rat neuronal culture labeled for microtubules and actin
Dr. Christophe Leterrier, NeuroCyto, INP, Marseille, France