Prof. Wendy Bickmore
Please tell us about your research.
I am interested in the three dimensional organization of the genome and understanding how that impacts how the genome functions, particularly with respect to gene expression. There are different ways of exploring that question, for example using some molecular tools. But I have always believed that light microscopy and imaging is a very powerful way to get insight into spatial organization. Fluorescence approaches are particularly great in this aspect.
Over the years we have explored different areas of genome organization: where chromosomes are located in the nucleus, where genes are located in the nucleus and the role of components in the nucleus that manage this organization. More recently we became interested in heterochromatin.
Heterochromatin is the inactive part of our genome, very often made up of repeated sequences from old retroviruses. Normally, heterochromatin is found in a rim around the edge of the nucleus. You can see it even by standard diffraction-limited light microscopy. But to see it much more cleanly and crisply one needs to use super-resolution imaging with a resolution of around 100 nm.
This organization is typical of most human cells, but there is a really interesting biological situation in which heterochromatin organization changes radically and that is called oncogene-induced senescence. Senescence is when cells stop dividing permanently. That can come about through a variety of factors, for examples as they age - through replicative senescence – cells just run out of steam and stop dividing. But they also do that when they are challenged with an activated oncogene such as Ras. This is thought to be an important tumor suppressor mechanism to stop tumors from developing.
What happens in human cells that are induced to go into senescence (because they are suddenly faced with an activated oncogene), is that the heterochromatin dramatically reorganizes away from the edge of the nucleus, and instead, moves into the middle of the nucleus into so-called senescence-associated heterochromatin foci.
This is really intriguing for someone who is interested in 3D genome organization - How you can have such a dramatic genome change and what causes it? In terms of the edge of the nucleus, most of my research and, I would say, most of the field has mostly focused on the proteins that are embedded in the nuclear membrane or just underneath it. We tended to ignore the fact that there are also holes in the nuclear membrane; that it is perforated by amazing macro molecular structures called nuclear pores. They are large structures in molecular terms, but very small in terms of cell biology and imaging. While looking for an explanation of why heterochromatin would lose its interaction with the periphery and move inside, we realized that something unusual was going on in the oncogene-induced senescence cell. They kept making the components of nuclear pores. Pores are extremely stable – once they are put into the nuclear membrane at the beginning of the cell cycle, they stay there for the rest of the cell cycle.
In normal situations, when cells stop dividing, they stop synthesizing the components of the nuclear pore. We realized however, that during oncogene-induced senescence, this does not happen. We wondered whether that change in the nuclear periphery has something to do with heterochromatin re-localization.
To address this question, we wanted to be able to count the nuclear pores. Since they are very tiny structures we used structured illumination microscopy, N-SIM, that gives us sufficient resolution in x and y. By using an antibody to detect nucleic pore components, we looked at the bottom surface of the nucleus and we were able to see a nice, resolved spot for each nuclear pore, all over the nuclear membrane. This allowed us to literally count nuclear pores on the images from N-SIM microscope in the cells before and after senescence.
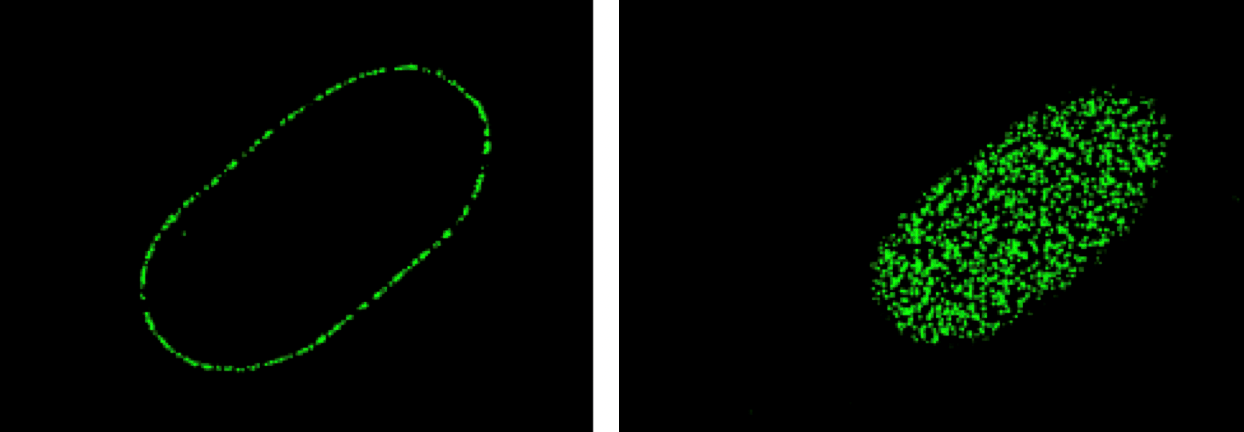
The left hand image shows a mid-plane section through the nucleus with pores visible around the rim of the nucleus.
The right hand image was taken of the bottom of the nucleus with pores embedded in the membrane.
What we found was consistent with the gene expression data – in oncogene-induced senescence the cells accumulated nuclear pores into the membranes. As a result, the portion of the nuclear membrane that contains these channels increases. What has been known for a long time in the field from electron microscopy images is that whereas heterochromatin is normally around the nuclear periphery, there is a gap wherever there is a nuclear pore. Our model was that it is the increased proportion of the nuclear periphery that is occupied by these heterochromatin-excluding pores that is responsible for kicking heterochromatin off the nuclear periphery and sending it to the middle of the nucleus. With this model in mind - that came from the N-SIM imaging - we were able to experimentally test it. Using siRNA we knocked down nuclear pore components and showed, again using the N-SIM, that that reduced the number of nuclear pores inserted in the nuclear membrane of the senescent cells and the internal foci heterochromatin had disappeared. Next, we wanted to know what component of the pore is important in this process. There are lots of different proteins in the pore; it is made up of different sub-structures. On the cytoplasmic side: there are cytoplasmic filaments and the core pore channel that sits in the membrane itself. On the nuclear side, which is the side we are interested in, there is the so-called a nuclear pore basket. This structure is thought to extend quite a way into the nucleus. The protein that sits on these nuclear pore baskets is called TPR. We reasoned that because TPR is closest to the chromatin in the nucleus, it might be the important protein responsible for brushing heterochromatin away from the pores. When we specifically knocked down TPR, but left the rest of the nuclear pore intact, we still had elevated numbers of nuclear pores in senescence, but they no longer had nuclear baskets with the TPR protein on them. We have found that that causes the internal foci heterochromatin to return to the nuclear periphery. This observation confirmed the hypothesis TPR keeps heterochromatin away from nuclear pores.
The N-SIM data helped us to reveal a brand new organizing principle for the nucleus which is exciting. But now we want to push it further.
Great, so what are your next steps and what challenges do you foresee?
We would like to know how far TPR filaments extend into the nucleus.
This will require turning to even higher resolution and we are thinking about labelling the extreme ends of the TPR protein which is predicted to be a very long filamentous protein, perhaps around 250 nm in size. We are considering using STORM to get precise localization on the N- and C-terminus of TPR to really look at the ultra-structure and the nano-structure around the pore and heterochromatin.
The great thing about the N-SIM is that you can apply it to standard biological samples - you do not need to change your methodology. You can use immunostaining that you would normally use for example in normal epi-fluorescence microscope.
I think that STORM is going to be more challenging. The labels become really critical in STORM microscopy and the sample preparation is much more challenging. That is the next thing that we are going to face and we will try to tackle it.
Also, we would definitely like to do live cell imaging of nuclear pore components, in particular of TPR and of heterochromatin itself. We are going to have to do some exploration of how much we can do super-resolution without killing the cells and the biological system. So that is another challenge ahead.
Sounds exciting! Please tell us a bit more about how find your experience using the N-SIM system?
We like N-SIM because of the applicability to our existing protocols. For an experiment we would normally have looked at on a wide-field microscope, we can just take the same sample and look at it on the N-SIM. We have worked hard to make sure that the quality of staining is good and we are really happy with the data that comes from the N-SIM and the reconstruction of that.
What my team particularly likes is also the software. They find that the interface really user-friendly.
What developments would you like to see for Super-resolution in the future?
What we would like in the future is to increase the speed of SIM. As mentioned before, cell-friendly super-resolution is needed. For microscopy developments more in general, I think that high-content imaging is going to be interesting. We would be really interested in imaging as a screening method, for example for small molecule inhibitors of processes. Being able to image lots of cells, lots of wells would be great. In my view, bridging the gap between light microscopy and electron microscopy, like correlative light electron microscopy is also going to be really exciting area to go into.
Great, thank you for your insights.
doi: 10.1101/gad.321117.118
