- zh Change Region
- Global Site
应用手册
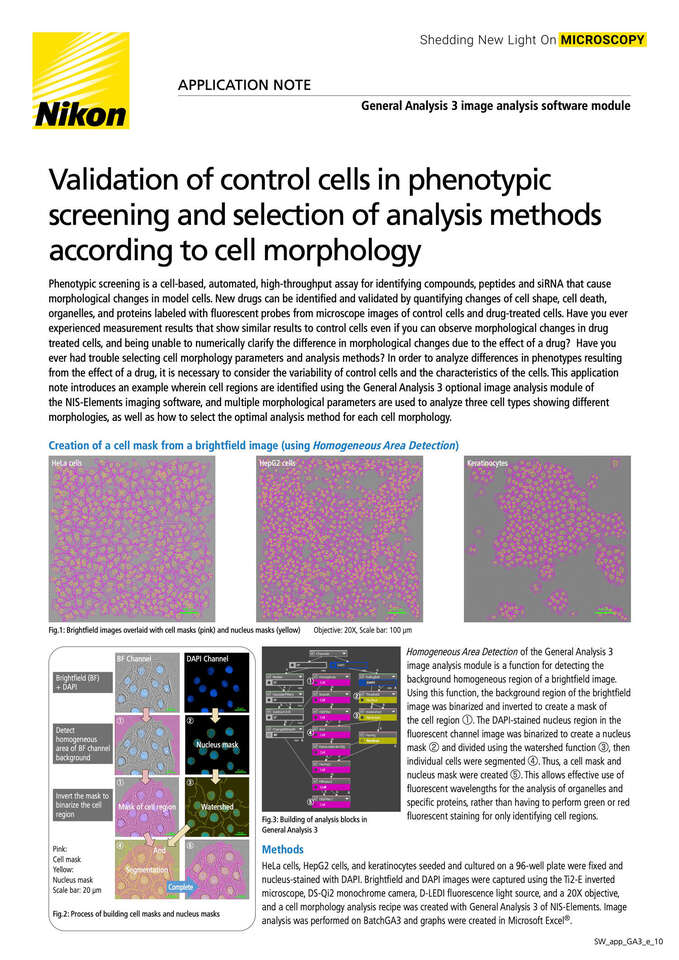
Validation of control cells in phenotypic screening and selection of analysis methods according to cell morphology
2022年3月
Phenotypic screening is a cell-based, automated, high-throughput assay for identifying compounds, peptides and siRNA that cause morphological changes in model cells. New drugs can be identified and validated by quantifying changes of cell shape, cell death, organelles, and proteins labeled with fluorescent probes from microscope images of control cells and drug-treated cells. Have you ever experienced measurement results that show similar results to control cells even if you can observe morphological changes in drug treated cells, and being unable to numerically clarify the difference in morphological changes due to the effect of a drug? Have you ever had trouble selecting cell morphology parameters and analysis methods? In order to analyze differences in phenotypes resulting from the effect of a drug, it is necessary to consider the variability of control cells and the characteristics of the cells. This application note introduces an example wherein cell regions are identified using the General Analysis 3 optional image analysis module of the NIS-Elements imaging software, and multiple morphological parameters are used to analyze three cell types showing different morphologies, as well as how to select the optimal analysis method for each cell morphology.
