- zh Change Region
- Global Site
应用手册
Label-free detection of neuron and astrocyte regions using deep learning technology
2022年10月
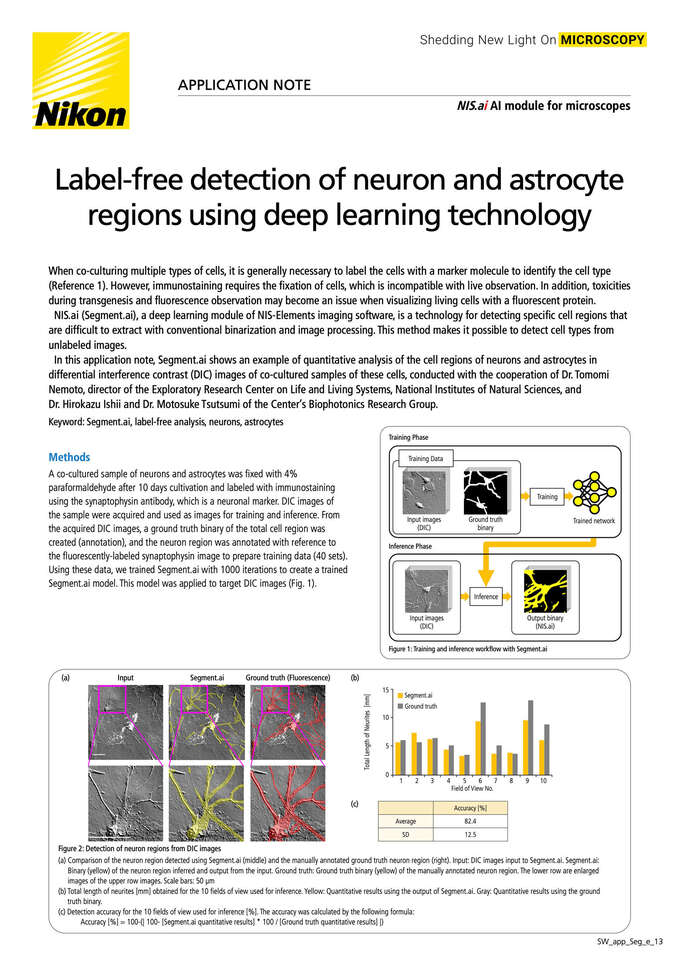
When co-culturing multiple types of cells, it is generally necessary to label the cells with a marker molecule to identify the cell type (Reference 1). However, immunostaining requires the fixation of cells, which is incompatible with live observation. In addition, toxicities during transgenesis and fluorescence observation may become an issue when visualizing living cells with a fluorescent protein.
NIS.ai (Segment.ai), a deep learning module of NIS-Elements imaging software, is a technology for detecting specific cell regions that are difficult to extract with conventional binarization and image processing. This method makes it possible to detect cell types from unlabeled images
In this application note, Segment.ai shows an example of quantitative analysis of the cell regions of neurons and astrocytes in differential interference contrast (DIC) images of co-cultured samples of these cells, conducted with the cooperation of Dr. Tomomi Nemoto, director of the Exploratory Research Center on Life and Living Systems, National Institutes of Natural Sciences, and Dr. Hirokazu Ishii and Dr. Motosuke Tsutsumi of the Center’s Biophotonics Research Group.
